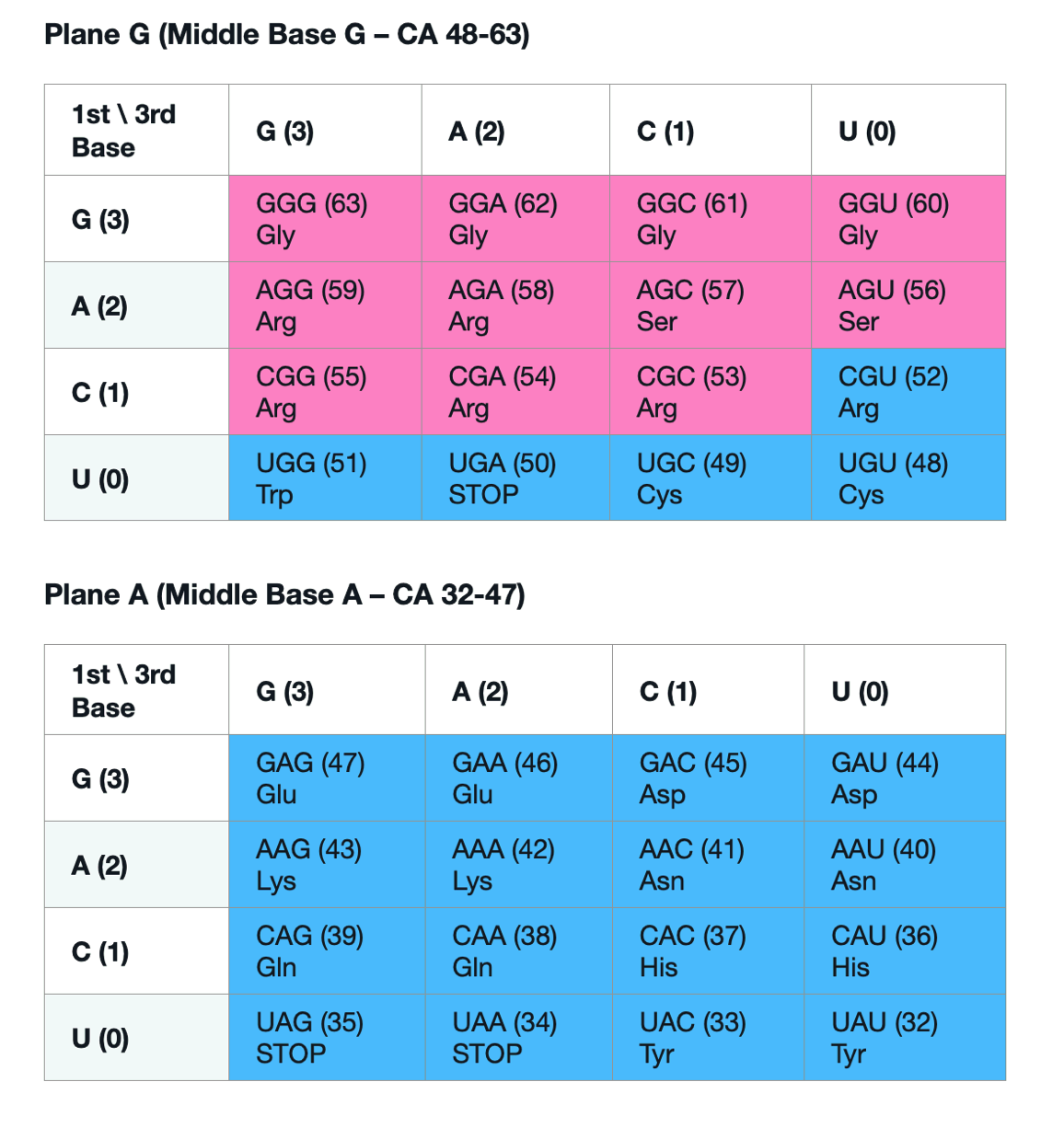
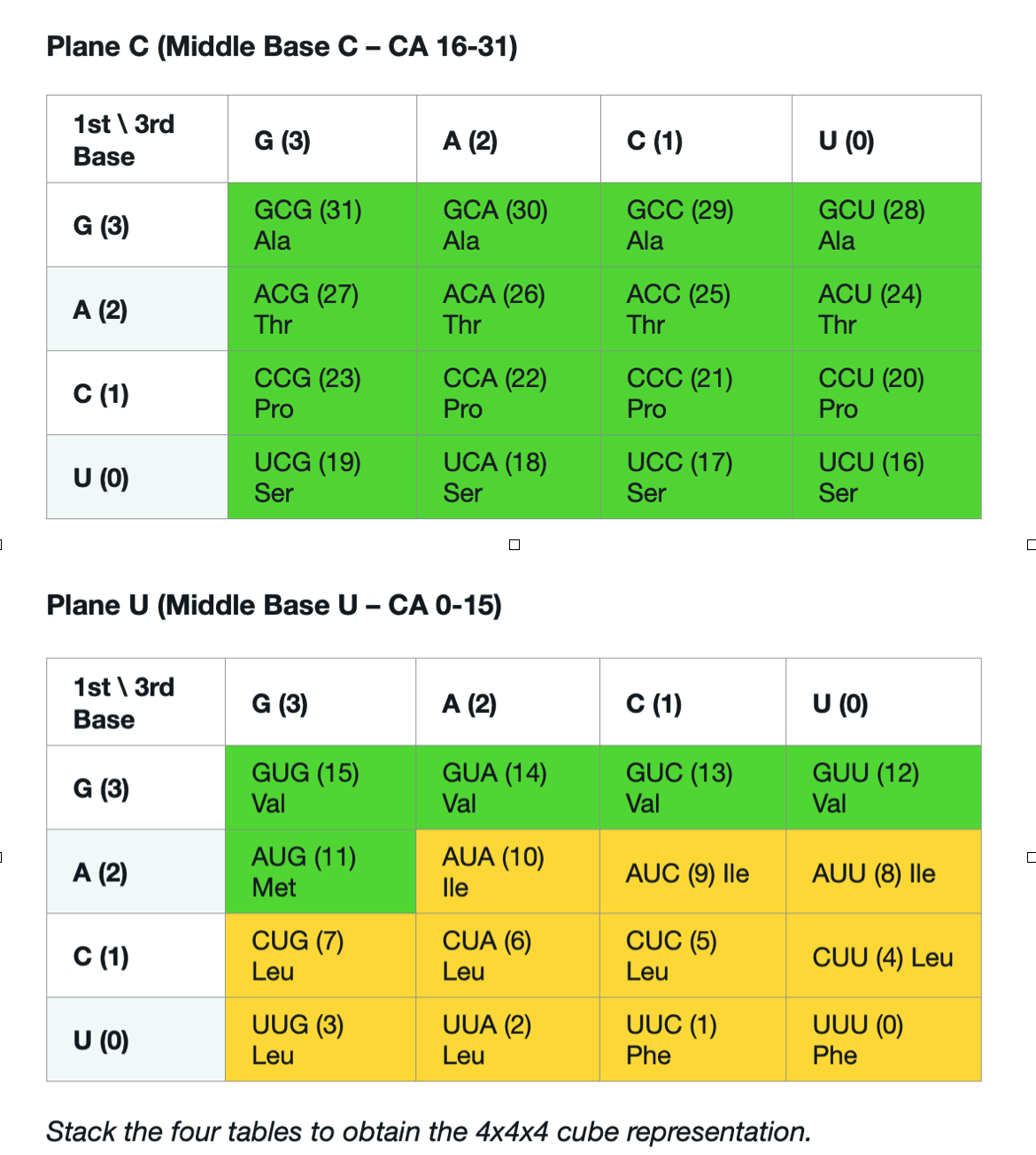
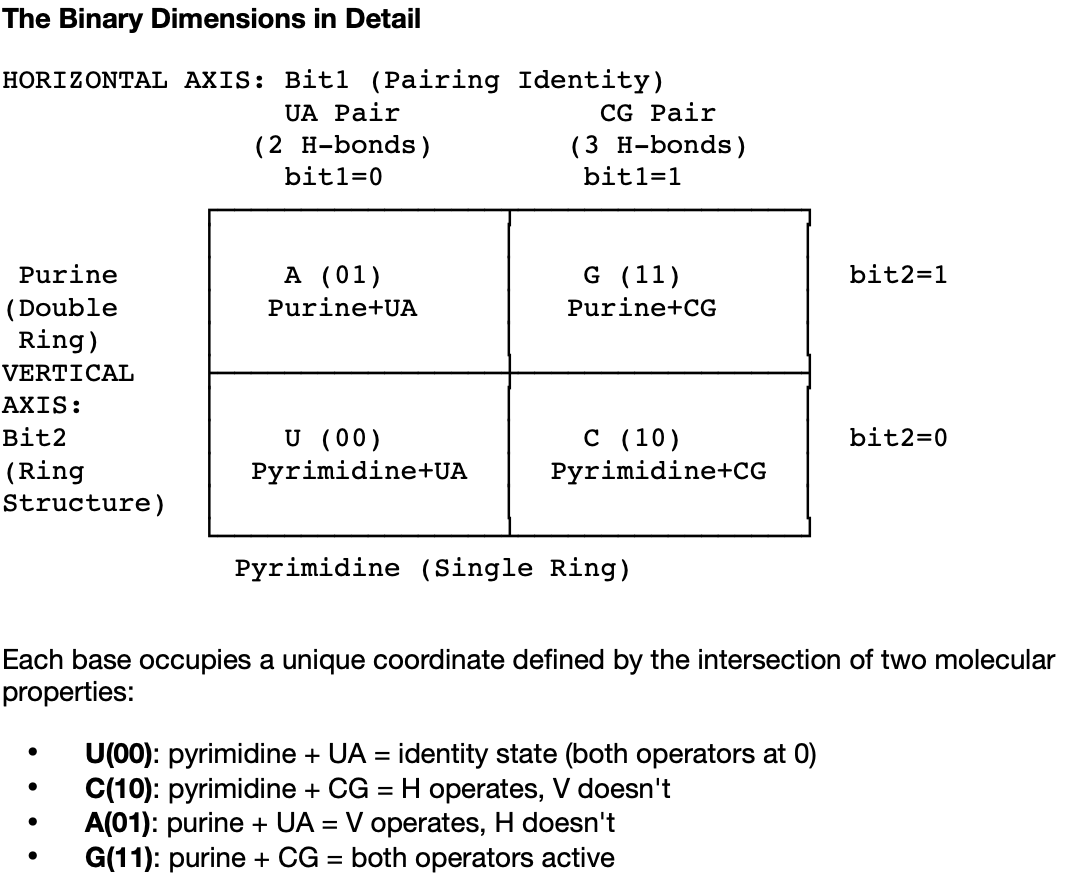
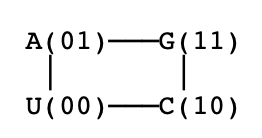From overclaim to conditional uniqueness
Paper 1. The 144 frameworks, filtered by the requirement that synonymous families stay contiguous. Two survive: UCAG [4,16,1] and its reverse reading GACU [4,16,1]. Fixing the reading direction by the ground state leaves UCAG [4,16,1] unique. The integer coordinate and center-dominant weighting are prior art (Sánchez 2005).
Paper 2. Structural consequences once UCAG [4,16,1] is fixed: the level structure, the divergence geometry, the graded mutation-step magnitudes. The ordering tracks wobble decoding geometry. These describe the chosen coordinate; they do not show the assignment was forced.
Paper 3. The 64-state address space is complete: positional properties give each codon a unique signature. This is a property of the addressing, and places no constraint on the degeneracy pattern or the assignment.
Paper 4. The general algebra fixes the weight set and the block-carrying position. Its stronger claim, that this position is forced to the center for odd n, does not hold without a layout assumption, which is why the center placement is sourced here from the degeneracy pattern instead.
Prior art. The integer coordinate and center-dominant weighting are Sánchez, Morgado and Grau (2005); the six-bit hypercube is Jiménez-Montaño and colleagues (1996); a separate uniqueness theorem for the assignment is Zamudio and José (2017). What this work adds is the substrate-based base order and the bounded-line serialization criterion, with leucine the deciding family. Verification against that corpus is pending.